
March 2021: Updates to PHO’s Variants of Concern Laboratory Testing
COVID-19
19 June 2021
Important note: We have made further updates to our variants of concern laboratory testing. Check out our new blog post: June 2021: Updates to COVID-19 Variants: Terminology, Laboratory Testing and Reporting to learn more.
Viruses are known to change and mutate, and SARS-CoV-2 (the virus that causes COVID-19) is no exception. As new and emerging variants become known, laboratory systems around the world, including our laboratory at Public Health Ontario (PHO) are required to adapt and shift to proactively monitor for these variants and support surveillance efforts to identify their spread.
How do we screen for VOCs at PHO?
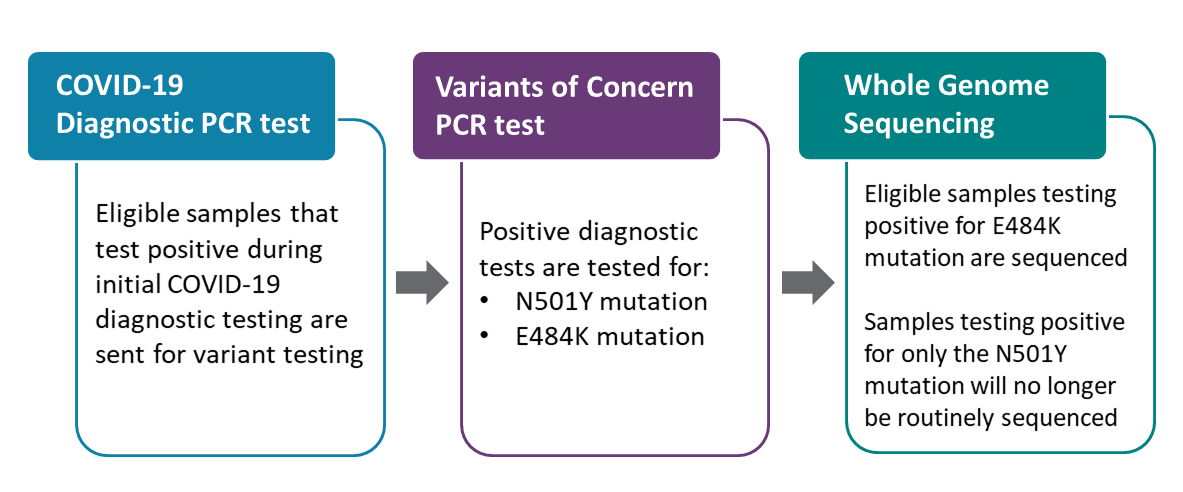
PHO’s laboratory has created a variant test that looks for mutations using real-time PCR (hereafter referred to as the PCR variant test). Eligible samples that tested positive during the initial COVID-19 diagnostic test are sent for PCR variant testing to determine if a mutation indicating a variant of concern (VOC) is present in the sample. Our test looked specifically for the N501Y mutation, which is the mutation found in the three predominant VOCs:
- B.1.1.7 (first identified in the UK)
- B.1.351 (first identified in South Africa)
- P.1 (first associated with Brazil)
Starting March 22, we have updated our PCR variant test to look for an additional mutation.
What has changed?
We have added a second target to our PCR variant test to also look for the E484K mutation, which is present in both B.1.351 and P.1. variants. Eligible COVID-19 samples that tested positive during initial diagnostic testing will now be tested for both the N501Y and E484K mutations at the same time. Eligible samples that test positive for the E484K mutation will be forwarded for genome sequencing. Samples that are N501Y positive and E484K negative are presumed to be likely B.1.1.7 based on current epidemiology and will no longer be routinely sequenced.
Why did we make this change?
PHO’s PCR variant test has been shown to be highly accurate in detecting the presence of the N501Y mutation. Given the accuracy of the PCR variant test, genome sequencing is no longer required to confirm that a sample is positive for a VOC. By introducing the E484K target in our PCR variant test, we will continue to track the B.1.1.7 variant and be able to more closely identify potential B.1.351, P.1 variants and other emerging variants.
What samples will go for whole genome sequencing?
Eligible samples that test positive for the E484K mutation will be forwarded for whole genome sequencing to help us better understand the epidemiology of emerging variants and how they are spreading in Ontario. Additionally, to support ongoing surveillance efforts, whole genome sequencing will continue to be conducted on a minimum of 5% of samples that do not have the E484K mutation.
Using PCR variant testing and genome sequencing in a targeted way ensures that labs have the capacity and resources to focus on the areas that will provide the most value in terms of detecting spread of B.1.1.7, B.1.351 and P.1 and in identifying new and emerging variants.
What is the difference between PCR variant and PCR diagnostic testing?
The overall goals of PCR variant testing and PCR diagnostic testing are different. The goal of PCR variant testing is for surveillance purposes and is intended to help us understand trends of the virus at a population level by testing the best quality samples. This is different from the goal of diagnostic testing, which is to diagnose people who are positive for the SARS-CoV-2 virus and for public health action to be taken to stop transmission of the virus.
If you are responsible for laboratory testing, check out our SARS-CoV-2 (COVID-19 Virus) Variant of Concern Surveillance Test Information Sheet and Labstract for more information.
Don’t have a MyPHO account? Register Now

