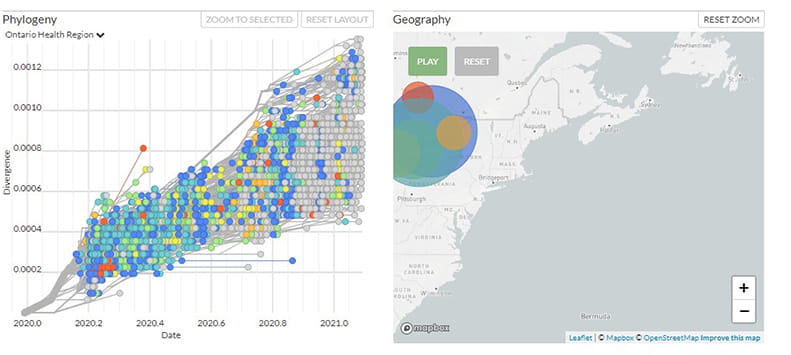
Phylogenetic Analysis of SARS-CoV-2 in Ontario
Genome sequencing, analysis and epidemiology of SARS-CoV-2 is important for understanding the evolution and spread of SARS-CoV-2 in Ontario. This interactive tool supports genomics researchers and experts, epidemiologists, virologists and public health experts explore the evolutionary relationships between SARS-CoV-2 sequences and view the genetic diversity across the entire genome.
Data in the tool can be filtered by:
Data in the tool can be filtered by:
- Ontario Health Region
- Pango lineages, including variants of concern lineages
- age group
- sex
- genotype
- country
- Clades: Nextstrain and GISAID
View Phylogenetic Analysis of SARS-CoV-2 in Ontario
The data in this tool includes SARS-CoV-2 samples sequenced at Public Health Ontario and does not include samples sequenced at other laboratories at this time.
The tool is updated Thursday between 4 and 6 p.m. ET and may be unavailable at this time. View the Technical Notes for more information on what is included in the tool.
The tool is best viewed using Google Chrome, Mozilla Firefox and Microsoft Edge.
Nextstrain
This interactive tool uses the open source platform Nextstrain, which provides powerful analytic and visualization tools to support genomic analysis of pathogens like SARS-CoV-2.
To learn more about Nextstrain or support using and interpreting data in the tool, visit the Nextstrain documentation webpage for a variety of resources, tutorials and user-guides.
Updated
3 Aug 2023
You need a MyPHO Account to save this page.
Log in to MyPHO
Don’t have a MyPHO account? Register Now
You have successfully created a MyPHO account!
Use MyPHO to save content relevant to you, take online courses and register for subscriptions.